Methods for Detecting Protein Phosphorylation
Introduction
Protein kinases transfer phosphate groups from ATP to serine, threonine, or tyrosine residues on protein peptide substrates, directly affecting the activity and function of the target. Radiolabel studies suggest that approximately 30% of proteins in eukaryotic cells are subject to phosphorylation.1, 2 This crucial post-translational modification regulates a broad range of cellular activities including the cell cycle, differentiation, metabolism, and neuronal communication. In addition, abnormal phosphorylation events are implicated in many disease states. When assessing phosphorylation, the method of choice may vary depending on many factors including the specific question being asked and availability of specialized equipment or reagents. This review provides a brief description of several methodologies currently used and addresses some of the benefits and drawbacks associated with each.
Table of Contents
Kinase Activity Assays
Protein kinases are often common elements in multiple signaling networks influencing numerous downstream effectors responsible for a biological response. Thus, assessing the activity of a single specific kinase may provide valuable information on parallel pathways.3 Kinase activity within a biological sample is commonly measured in vitro by incubating the immunoprecipitated kinase with an exogenous substrate in the presence of ATP. Measurement of the phosphorylated substrate by a specific kinase can be assessed by several reporter systems including colorimetric, radioactive, or fluorometric detection.4 Additionally, R&D Systems offers a non-radioactive Universal Kinase Activity Kit that allows for the quantification of kinase activity for any kinase that produces ADP. Although information can be obtained regarding the actions of a specific kinase, assessing enzyme activity in cellular extracts only provides a glimpse of the signaling landscape. Little is revealed about the proteins being modified, and in vitro activity assays do not address the role of potential endogenous phosphatase activity. Direct detection of phosphorylated proteins can provide a more detailed analysis of the cellular response to an external stimulus, as identification of a phosphopeptide provides information regarding the expression and the functional state of that protein.
Phospho-Specific Antibody Development
A classical method of directly measuring protein phosphorylation involves the incubation of whole cells with radiolabeled 32P-orthophosphate, the generation of cellular extracts, separation of proteins by SDS-PAGE, and exposure to film.2, 5 This labor-intensive method requires many multi-hour incubations and the use of radioisotopes. Other traditional methods include 2-dimensional gel electrophoresis, a technique that assumes phosphorylation will alter the mobility and isoelectric point of the protein.
In light of these laborious methods, the development of phosphorylation-dependent antibodies was a welcome event for researchers. In 1981, the first documented phospho-antibody was produced in rabbits immunized with benzonyl phosphonate conjugated to keyhole limpet hemocyanin (KLH). This antibody broadly recognized proteins containing phosphotyrosine.6 Ten years later, phosphorlyation state-specific (phospho-specific) antibodies were developed by immunizing rabbits with synthetic phosphopeptides representing the amino acid sequence surrounding the phosphorylation site of the target protein.7 The immune sera was applied to a peptide affinity column to generate a highly specific immunoreagent. The availability of phospho-specific antibodies has opened the door for the improvement of traditional methods as well as the development of new immunoassay techniques. The main caveat in utilizing phospho-specific antibodies in any technique is that successful detection is dependent on the specificity and affinity of the antibody for the phospho-protein of interest.
Simple Western Capillary Western Assays
Simple Western™ is a next-generation capillary immunoassay technology that performs automated Western blot analysis inside a benchtop instrument like Jess™, Abby™, Peggy Sue™, Sally Sue™, and NanoPro™ 1000. Compared to traditional methods like Western blot, Simple Western instruments can detect and quantify phosphorylated isoforms of target proteins with high specificity, reproducibility, and sensitivity. The Stellar NIR/IR Modules for Jess lead the industry in fluorescence detection sensitivity for Western blotting workflows, with 100X greater sensitivity than the closest competing fluorescence imaging technology. As an open immunoassay platform using conventional antibodies for detection, Simple Western assays can detect phosphorylated proteins in two ways:
1. Charge-Based Simple Western Assays
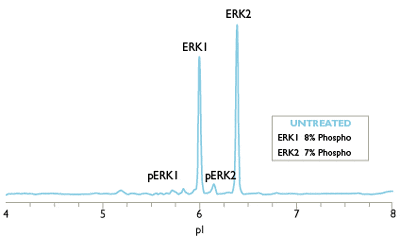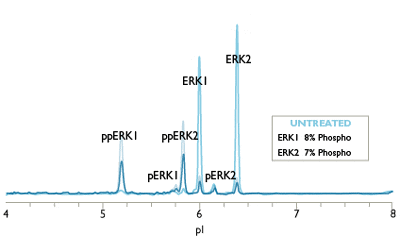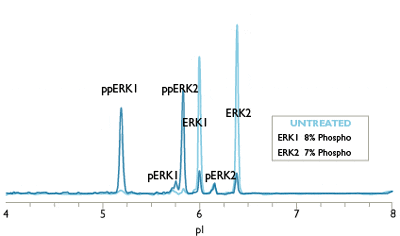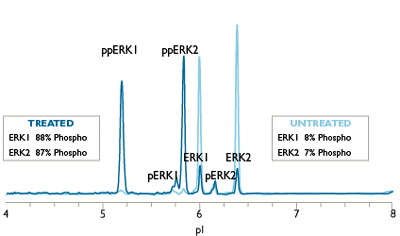
Charge-based Simple Western assays separate protein samples by isoelectric point (charge-based separation) to resolve all phosphorylated isoforms of your target protein.
For example, monophospho- and diphospho-ERK isoforms are not resolved by traditional Western blot analysis, and the sample quantity required is relatively large. The Simple Western ERK1/ERK2 assay can distinguish and quantify unphosphorylated, mono, and dual-phosphorylated isoforms of ERK1 and ERK2, allowing a more accurate determination of ERK activation than Western blots provide (Figure 1). Phosphatase treatment may be used to further validate phospho-protein isoforms on Simple Western.
2. Size-Based Simple Western Assays
Size-based Simple Western assays separate protein samples by molecular weight (size separation) using phospho-specific antibodies for detection.
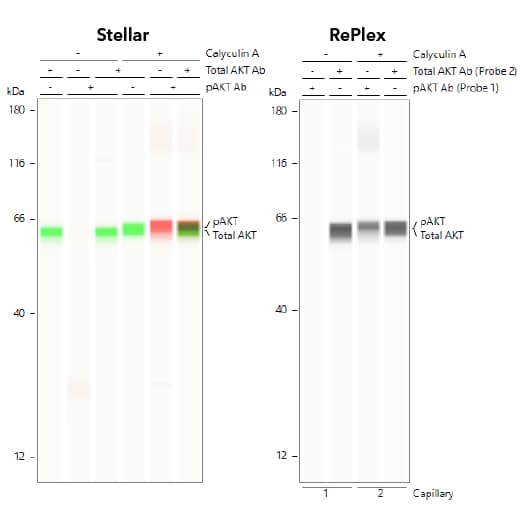
In addition, Simple Western offers flexible multiplex strategies with sequential multi-channel detection to characterize phospho-protein and total-protein isoforms in the same sample. For example, Stellar assays on Jess provide high-sensitivity multiplexing in NIR/IR fluorescence and chemiluminescence channels (Figure 2, left panel). RePlex assays on Jess and Abby perform two sequential cycles of immunodetection by removing the antibodies from the first probing cycle for a second probing cycle using fresh antibodies (Figure 2, right panel). Unlike the cumbersome membrane strip and reprobe method of Western blots, RePlex assays are not subject to interference of sample loss between probing cycles because samples are covalently immobilized to the capillary walls and conditions are optimized for efficient removal of antibodies. Furthermore, Simple Western can perform simultaneous total protein detection which can be used to normalize your data for more accurate protein expression results.

Western Blot
The Western blot is the most common method used for assessing the phosphorylation state of a protein, and most cell biology laboratories possess the equipment necessary to perform these experiments. Following separation of the biological sample with SDS-PAGE and subsequent transfer to a membrane (usually PVDF or nitrocellulose), a phospho-specific antibody can be used to identify the protein of interest (View phospho-specific antibodies for Western blot analysis). The typical Western blot protocol eliminates the hazards and waste disposal requirements associated with the use of radioisotopes.
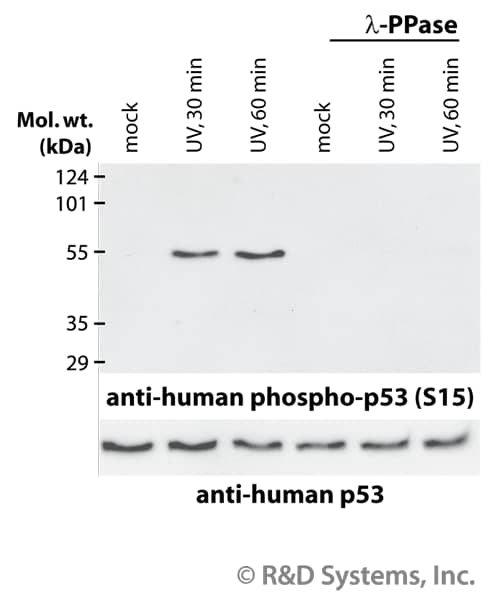
Many phospho-specific antibodies are quite sensitive and can readily detect the phosphorylated protein in a routine sample (e.g., 10-30 µg whole cell extract). Because the measured levels of a phospho-protein may change with treatment or through gel loading errors, researchers often utilize an antibody that detects the total level of the cognate protein (regardless of phosphorylation state) to determine the phosphorylated fraction relative to the total fraction and to serve as an internal loading control (Figure 3). Both chemiluminescent and colorimetric detection methods are common, and molecular weight markers are also generally used to provide information about protein mass. A detailed Western blot protocol can be found at R&D Systems.
Phosphorylated p53 in CEM Cells
Figure 3. Human T lymphoblast CEM cells were exposed to UV-C light. Cellular extracts generated at 30 or 60 minutes post-irradiation were assessed by Western blot using rabbit anti-human phospho-p53 (S15) polyclonal antibody (Catalog # AF1043, upper panel) or goat anti-human p53 polyclonal antibody (Catalog # AF1355, lower panel). Indicated samples were treated with lambda-phosphatase (lambda-PPase).
Enzyme-Linked Immunosorbent Assay (ELISA)
The ELISA has become a powerful method for measuring protein phosphorylation. ELISAs are more quantitative than Western blotting and show great utility in studies that modulate kinase activity and function. The format for this microplate-based assay typically utilizes a capture antibody specific for the desired protein, independent of the phosphorylation state. The target protein, either purified or as a component in a complex heterogeneous sample such as a cell lysate, is then bound to the antibody-coated plate. A detection antibody specific for the phosphorylation site to be analyzed is then added. These assays are typically designed using colorimetric or fluorometric detection. The intensity of the resulting signal is directly proportional to the concentration of phosphorylated protein present in the original sample (Figure 4). The phospho-specific ELISA technique confers several advantages over more traditional immunoblotting in the measurement of protein phosphorylation. First, results are easily quantifiable by utilizing a calibrated standard. Second, high specificity is possible due to the use of two antibodies specific for the target protein employed together in the sandwich format. Thirdly, the higher sensitivity often accomplished using ELISAs allows for smaller sample volumes and the detection of low abundance proteins. Finally, the microplate-based format allows for much higher throughput than traditional Western blotting (View Surveyor IC ELISA Kits). ELISAs generally provide an indirect measurement of kinase activity. However, variations in the technique described above use an immobilized capture antibody, substrate, and a phospho-substrate detection method for more direct measurements of kinase activity.
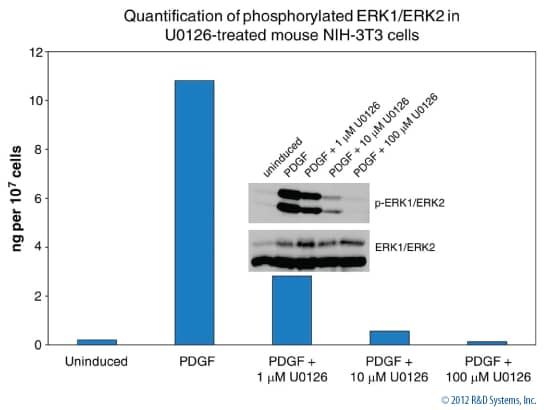
Phosphorylated ERK1/ERK2 in NIH-3T3 Cells
Figure 4. NIH-3T3 cells were treated with 100 ng/mL of human PDGF (Catalog # 120-HD) for 10 minutes in the presence or absence of the MEK1/2 inhibitor U0126 (Catalog # 1144). Following cell lysis, phosphorylated ERK1 and ERK2 were quantified with the Surveyor IC Immunoassay kit (Catalog # SUV1018B). The results are highly comparable between the Surveyor IC Immunoassay results and the amounts of phosphorylated ERK1/ERK2 (Catalog # AF1018) relative to total ERK1/ERK2 (Catalog # MAB1576) detected by Western blot (inset).
Cell-Based ELISA
Although in vitro biochemical kinase assays such as the typical sandwich ELISA are routinely used for hypothesis testing and drug screening, they cannot replicate the intracellular environment. Analyzing protein phosphorylation within intact cells may more accurately represent the status of specific signaling networks. Several immunoassays enabling the measurement of protein phosphorylation in the context of a whole cell have recently been developed. The cells are stimulated, fixed, and blocked in the same well. Phospho-specific antibodies are used to assess phosphorylation status using fluorometric (Figure 5) or colorimetric detection systems. Furthermore, the phospho-protein and total protein are simultaneously detected in the same microplate well. Therefore, signals derived from the target protein can be normalized to that of the second protein, correcting for well-to-well variations and allowing phospho-protein levels to be accurately assessed and compared across multiple samples, similar to using phospho-specific and total protein antibodies in a traditional immunoblot. These assays bypass the need for the creation of cell lysates and are therefore more amenable to high throughput analyses (View Cell-Based ELISA Kits).
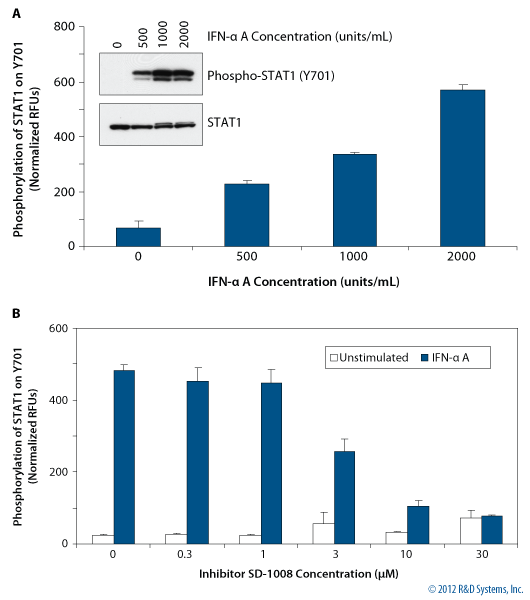
Measurement of STAT1 (Y701) Phosphorylation in HeLa Cells
Figure 5. (A) HeLa cells were treated with human IFN-alpha A (Catalog # 11100-1) for 20 minutes. After fixation of cells, phosphorylation of STAT1 (Y701) was determined and normalized to total STAT1 in the same well using the Phospho-STAT1 (Y701) Cell-Based ELISA kit (Catalog # KCB2894). Values represent mean + range of duplicate determinations. Analysis of phosphorylated STAT1 (Catalog # AF2894) and total STAT1 (Catalog # MAB14901) by Western blotting is also shown (inset A). (B) HeLa cells were pretreated for 30 minutes with the Janus kinase 2 (JAK2) inhibitor SD-1008 (Catalog # 3035) and then either left unstimulated or treated with 2000 units/mL IFN-alpha A for 20 minutes. Phosphorylation of STAT1 (Y701) was determined as in (A).
Intracellular Flow Cytometry and ICC/IHC
The traditional techniques of intracellular flow cytometry and immunocytochemistry/immunohistochemistry (ICC/IHC) are powerful tools for detecting phosphorylation events.8, 9 Flow cytometry uses a laser to excite the fluorochrome used for antibody detection. Filter sets and fluorochromes with non-overlapping spectra must be carefully chosen when assessing multiple proteins in the same cell. Flow cytometry is advantageous because it allows for rapid, quantitative, single cell analysis (Figure 6).10 Proteins can be detected in a specific cell type within a heterogeneous population via cell surface marker phenotyping without the need to physically separate the cells. In this way, a small, rare population of cells may be analyzed without concern for cell loss or altered protein expression that may occur during a cell-sorting process (View phospho-specific antibodies for flow cytometry).
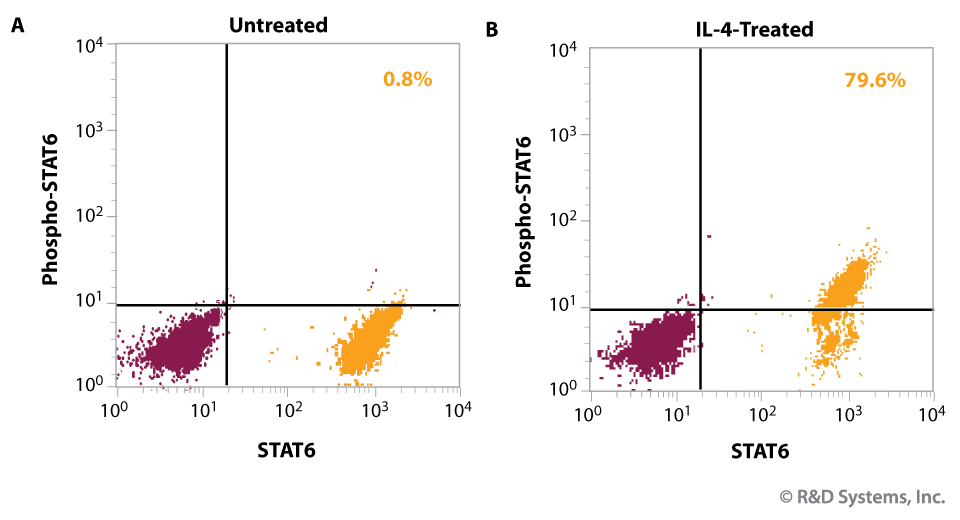
Detection of IL-4-induced STAT6 Phosphorylation by Intracellular Flow Cytometry
Figure 6. Human Daudi lymphoblastoid cells, (A) untreated or (B) activated with recombinant human IL-4 (Catalog # 204-IL) were fixed and permeabilized with methanol. Simultaneous detection (orange) of total (x axis) and phosphorylated (y axis) STAT6 was performed by co-staining cells with allophycocyanin-conjugated anti-STAT6 (Catalog # IC2167A) and phycoerythrin-conjugated anti-phospho-STAT6 (Catalog # IC3717P), respectively. The percent of cells positive for phospho-STAT6 is indicated in each treatment (upper right). Staining with isotype controls (Catalog # IC003A and # IC105P; maroon) highlights the specificity of the STAT6 antibodies.
ICC generally refers to protein detection by microscopy in cultured cells, while IHC refers to protein detection in intact tissue sections. Like flow cytometry, these techniques allow for the assessment of multiple proteins within a cell or tissue provided that adequate attention is given to avoid overlapping fluorescence spectra or color. Both fluorescent and colorimetric detection techniques are commonly used (Figure 7). In contrast to other formats for monitoring phosphorylation, ICC is usually the method of choice for determining intracellular localization (View phospho-specific antibodies for ICC/IHC). Both flow cytometry and ICC/IHC require high-affinity and high-specificity antibodies, blocking steps, controls, and antibody titration to eliminate ambiguous results due to non-specific binding.
Detection of phospho-proteins by flow cytometry and ICC require that the protein is stable and accessible to the antibody. Cells are usually stimulated and fixed with formaldehyde or paraformaldehyde to cross-link the phospho-proteins and stabilize them for analysis. The fixed cells must then be permeabilized to allow for entry of phospho-specific antibodies into the cells. Different permeabilization techniques are often useful for various subcellular locations. A mild detergent will allow for detection of cytoplasmic proteins, while alcohol may be required for antibody access to nuclear proteins. Alcohol permeabilization may also enhance phospho-protein detection using peptide specific antibodies due to the denaturing property of alcohol.10 For more detailed ICC/IHC protocols, please visit our website at www.rndsystems.com/protocol-types/iccihc-protocols.

Detection of Phosphorylated Proteins Using ICC/IHC
Figure 7. Human Daudi lymphoblastoid cells were treated with (A) recombinant human IL-4 (Catalog # 204-IL) or (B) left untreated. Phosphorylated STAT6 was detected using anti-human phospho-STAT6 polyclonal antibody (Catalog # AF3717), followed by staining with NorthernLights™ 557-conjugated goat anti-rabbit IgG (Catalog # NL004; red) and DAPI nuclear staining (blue). C: Phosphorylated ERK1/ERK2 was detected in a section of inflamed rat brain cortex using anti-human/mouse/rat phospho-ERK1/ERK2 polyclonal antibody (Catalog # AF1018). The tissue was stained with the anti-rabbit HRP-DAB Cell and Tissue Staining Kit (Catalog # CTS005; brown) and counterstained with haematoxylin (blue).
Mass Spectrometry
A comprehensive assessment of protein phosphorylation (phosphoproteomics) in complex biological samples, such as cell lysates, is important for understanding phosphorylation-based signaling networks. Large-scale phospho-protein analysis in complex protein mixtures involves identification of phospho-proteins and phosphopeptides and sequencing of the phosphorylated residues. Mass spectrometry (MS) techniques are useful tools for these tasks. Although MS can be used with excellent sensitivity and resolution to identify a single protein, there are several inherent difficulties for the analysis of phospho-proteins. First, signals from phosphopeptides are generally weaker, as they are negatively charged and poorly ionized by electrospray MS, which is performed in the positive mode.11 Second, it can be difficult to observe the signals from low-abundance phospho-proteins of interest in the high-background of abundant non-phosphorylated proteins. To overcome these drawbacks, several enrichment strategies for phospho-protein analysis by MS have been developed including immobilized metal affinity chromatography (IMAC),12 phosphospecific antibody enrichment,13 chemical-modification-based methods such as beta-elimination of phospho-serine and -threonine,14 and replacement of the phosphate group with biotinylated moieties.15
Multi-Analyte Profiling
Mass spectrometric techniques such as collision-induced dissociation (CID) and electron transfer dissociation (ETD) provide comprehensive parallel analysis of peptide sequences and post-translational modifications such as phosphorylation.16 These techniques are labor-intensive, and strategies for comprehensive phosphorylation analysis may not be needed if particular pathways are of primary interest. This has led to the development of several novel methods for measuring protein phosphorylation of multiple analytes simultaneously. In general, these involve the use of phospho-specific antibodies and include microplate-based (View Proteome Profiler 96 Antibody Arrays) and membrane-based (View Proteome Profiler Antibody Arrays) detection formats (Figure 8-12).17,18 The obvious benefit of these assays is that throughput capability is greatly enhanced by bypassing the need for running multiple individual Western blots or traditional ELISA-based assays. These techniques are also known for providing more data while requiring very little sample volume. In trade, protein profiling assays are typically recognized as being less sensitive than their more conventional counterparts due to potential antibody cross-reactivity.
Phospho-Protein Multiplex Assays
Our Phospho-specific Antibody Arrays Dominate Primary Literature
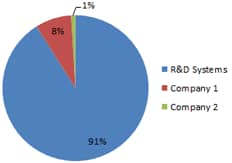
Figure 8. Phospho-specific Antibody Arrays Referenced in Primary Literature. There are over 200 references to phospho-specific, membrane-based antibody arrays in the primary literature. Proteome Profiler™ Phospho-specific Antibody Arraysoffered by R&D Systems account for 91% of those referenced.

Data from Our Phospho Antibody Arrays Are Published in Journals
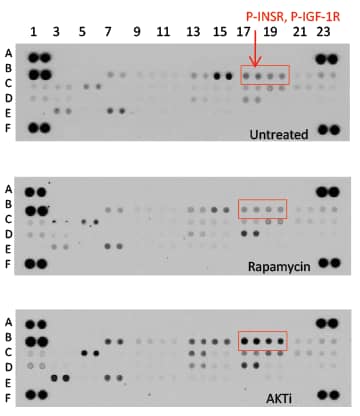
Figure 9. Proteome Profiler Phospho-RTK Antibody Arrays were Utilized to Investigate RTK Activation in Breast Cancer Cells.
Fold induction P-RTK in 8T-474 |
||
| P-RTK | AKTi | Rapa |
| ErbB3 | 12.1 | 4.6 |
| ErbB4 | 3.2 | 1 |
| INS R | 2.4 | 0.5 |
| IGF-I R | 2.3 | 0.6 |
| Flt-3 | 6.1 | 1.2 |
| MSP R | 2.9 | 1.5 |
| Mer | 1.7 | 2 |
| EphA1 | 2.4 | 0.7 |
| EphA7 | 3.1 | 0.5 |
| MuSK | 1.3 | 2 |
| MET | 0.2 | 1 |
Table from Chandarlapaty et al.5

New Microplate-based Proteome Profiler 96 Phospho Antibody Arrays
Features
- Up to 16 analytes/well
- Quick 3 hour assay time
- Small sample size requirement
- Chemiluminescent and near-infrared detection
- Amenable to high throughput
Figure 10. Analysis of RTK Phosphorylation. The MDA-MB-453 human breast cancer cell line was treated with NRG1-beta 1 (Catalog # 396-HB). Phosphorylation of 8 receptor tyrosine kinases (RTKs) was assessed in cell lysates using the (A) Proteome Profiler 96 Human Phospho-RTK Antibody Array 2, Infrared (Catalog # ARZ002NIR) or the (B) Proteome Profiler 96 Human Phospho-RTK Antibody Array 2 (Catalog # ARZ002). (C) User-defined Proteome Profiler 96Custom Phospho Arrays** (learn more) are also available.
Detection of MAP Kinase Phosphorylation
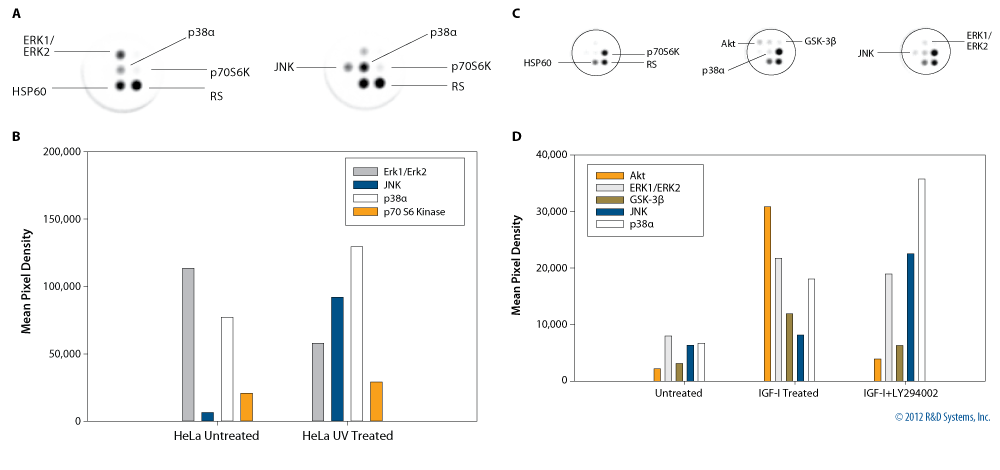
Figure 11. Lysates from HeLa cells (20 mg/well) that were either untreated or exposed to 150 J/m2 of UV light followed by a 30 minute recovery period were tested using the Proteome Profiler 96 Human Phospho-Kinase Array 1 Kit (Catalog # ARZ004). Array signals were analyzed using Q-View image analysis software. Inverted images of the wells are shown (A) with mean pixel densities in vertical bar graphs (B). MCF-7 cells were either untreated or treated with 100 ng/mL of human IGF-I (Catalog # 291-G1) for 1 hour; cells that received IGF-I were either pre-treated or not with the PI3K inhibitor LY294002 (Catalog # 1130) (C). Array signals were analyzed using Q-View image analysis software. Inverted images of the wells are shown (C) with mean pixel densities in vertical bar graphs (D). (RS = Reference Spot).
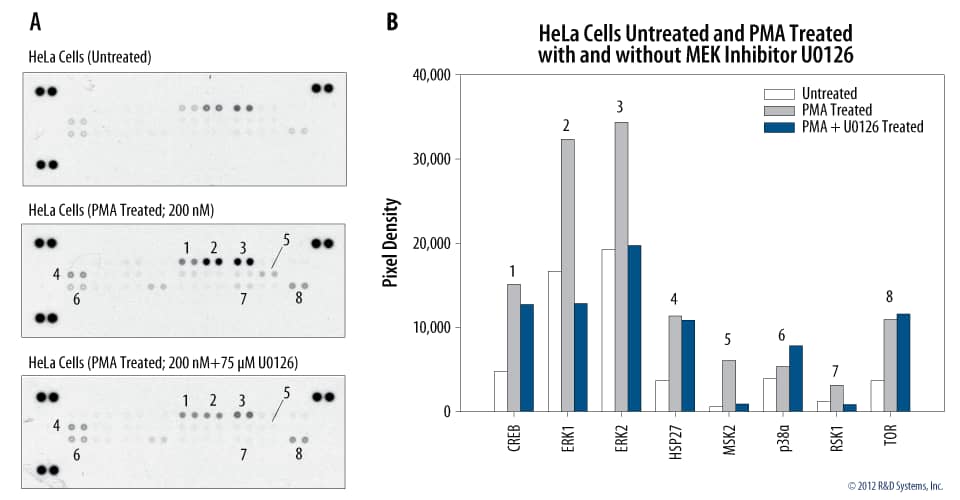
The Effect of the MEK1/2 Inhibitor U0126 on the ERK Pathway.
Figure 12. (A) Proteome Profiler Human Phospho-MAPK Array (Catalog # ARY002B) was incubated with lysates from HeLa cells untreated or treated with PMA in the presence or absence of U0126 (Catalog # 1144). (B) Graphical representation of the data.
Conclusion
Assessing protein phosphorylation is often an essential component of the cell biologist's repertoire for understanding intracellular factors underlying cellular activities. Given the important role kinases play, it is critical for researchers to have quality tools for measuring protein phosphorylation and/or kinase activity. Each technique excels in different contexts, and care must be taken to choose the method that best fits the experimental design. This review provides a brief glimpse of several of the most widely used methods for assessing protein phosphorylation. Because of a growing demand, methodologies continue to improve, bringing researchers closer to understanding these complex and important processes that ultimately control cellular function.
References
- Sefton, B.M. & S. Shenolikar (1996) Current Protocols in Molecular Biology 18.1.1-18.1.5.
- Ubersax, J.A. & J.E. Ferrell (2007) Nat. Rev. Mol. Cell Biol. 8:530.
- Ni, Q. et al. (2006) Methods 40:280.
- Johnson, S.A. & T. Hunter (2005) Nat. Methods 2:17.
- de Graauw, M. et al. (2006) Electrophoresis 27:2676.
- Ross, A.H. et al. (1981) Nature 294:654.
- Czernik, A.J. et al. (1991) Methods Enzymol. 201:264.
- Zell, T. et al. (2001) Proc. Natl. Acad. Sci. USA 98:10805.
- Willinger, T. et al. (2005) J. Immunol. 175:5895.
- Krutzik, P.O. et al. (2004) Clin. Immunol. 110:206.
- Mann, M. et al. (2002) Trends Biotechnol. 20:261.
- Brill, L.M. et al. (2004) Anal. Chem. 76:2763.
- Steen, H. et al. (2002) J. Biol. Chem. 277:1031.
- Zhou, H. et al. (2001) Nat. Biotechnol. 19:375.
- Oda, Y. et al. (2001) Nat. Biotechnol. 19:379.
- Molina, H. et al. (2007) Proc. Natl. Acad. Sci. USA 104:2199.
- Stiehl, D.P. et al. (2012) Oncogene 31:2283.
- Bai, Y. et al. (2012) Cancer Res. 72:2501.