Confidently Visualize Oncology Biomarkers in Tissues and Make Meaningful Discoveries
"RNAscope is a robust and reproducible platform that performs reliably across a wide range of clinical specimens. It has been instrumental in validating key biomarkers, defining molecular subtypes, and supporting several of our peer-reviewed publications. The technology continues to be an essential component of our spatial molecular pathology and biomarker discovery."
- Nallasivam Palanisamy, MSc., MPhIL., PhD, Associate Professor, Henry Ford Cancer Institute
Confidently Visualize Oncology Biomarkers in Tissues and Make Meaningful Discoveries
RNAscope™ brings oncology biomarkers into spatial context. It shows exactly where signals sit within intact tumor tissue while preserving morphology. At single-cell resolution, you can define cell types, interactions, and tumor architecture directly in situ.
With multiomics, RNAscope pairs with protein detection to visualize RNA and protein in the same section. This connects molecular signals to the tumor microenvironment with high specificity.
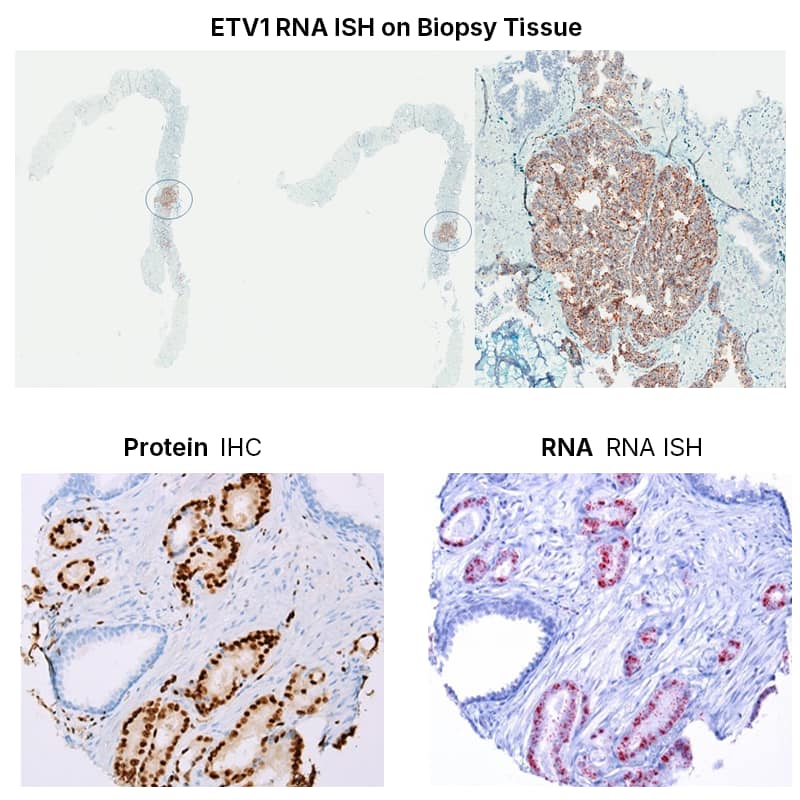
Customer highlight example to the right shows multiplex RNAscope with immunohistochemistry mapping ETS rearrangements and tumor heterogeneity with strong signal, low background, and clear tissue context.
Nallasivam Palanisamy, MSc., MPhIL., PhD Associate Professor, Henry Ford Cancer Institute
In our laboratory, we have extensively implemented the RNAscope- RNA in situ hybridization platform from Advanced Cell Diagnostics/Bio-Techne as part of our translational research program in prostate cancer. Our work focuses on understanding molecular subtypes and tumor heterogeneity, and RNAscope has provided a highly sensitive and spatially resolved method for detecting clinically relevant RNA transcripts in formalin-fixed, paraffin-embedded (FFPE) tissues.
We have successfully applied multiplex RNAscope assays to detect ETS family gene rearrangement–associated transcripts, including ERG, ETV1, ETV4, and ETV5, within the same tissue sections. This approach was further integrated with immunohistochemistry for ERG and SPINK1, allowing simultaneous evaluation of RNA and protein expression in the native histologic context. The combined RNA–protein platform enabled us to characterize both intertumoral and intratumoral molecular heterogeneity with high precision.
RNAscope consistently produced strong, punctate signals with excellent signal-to-noise ratios, even for low-abundance transcripts. The assay’s probe design and amplification chemistry provided high specificity, which was critical for distinguishing closely related ETS family members and for accurately mapping molecularly distinct tumor foci. The method also preserved tissue morphology, facilitating direct correlation with histopathologic features.
In our experience, RNAscope is a robust and reproducible platform that performs reliably across a wide range of clinical specimens. It has been instrumental in validating key biomarkers, defining molecular subtypes, and supporting several of our peer-reviewed publications. The technology continues to be an essential component of our spatial molecular pathology and biomarker discovery workflows, particularly for studies focused on tumor heterogeneity and clinically actionable molecular alterations.
Upcoming New Webinar
06/17/2026 / 8:00 AM PT / Resolve Oncology Biomarkers and Spatially Map Diversity in Prostate Cancer TME
To learn more about how RNAscope spatial multiomics solutions are used in oncology applications to identify cancer biomarkers, study splice variants, point mutations, interrogate TME, advance cell therapy and antibody therapeutic programs visit our oncology page.