Exosome Characterization with Simple Western
Introduction
Exosomes are small endosome-derived lipid nanoparticles approximately 30-200 nm in diameter that are actively secreted by exocytosis by most living cells.1 Exosome release can occur constitutively or upon induction, under pathological or non-pathological conditions, and in a dynamic, regulated and functionally-relevant manner. The molecular composition and quantity of released exosomes depend on the state of the parent cell. Exosomes are isolated from diverse cell lines and primary cultures, as well as from biological fluids like serum, plasma and saliva. Given this diversity, exosomes have pleiotropic physiological functions, and it is becoming increasingly clear that they have an important role in many pathological conditions such as cancer, infectious diseases, and neurodegenerative diseases. Hence, exosomes are a prime target in liquid biopsy methods.2 Exosomes may also be engineered as promising vesicles for drug delivery.3
Due to their tiny size, isolating exosomes from complex biological matrices can be a challenge, and yields are often low. Here, Simple Western™ is an ideal solution for exosome characterization because it requires only 3 µL of sample and offers picogram-level sensitivity, reducing the number of exosomes typically required for traditional Western blot. Exosome characterization can be performed using appropriate detection antibodies against exosome-associated antigens. These target antigens may be generic exosome markers or specific to the cell or fluid sample from which they originate. In this protocol, we use Simple Western to examine both generic and sample dependent targets using commercially-available exosome standards and antibodies.
Materials
- Instrument: Jess™, Abby™, Wes™, Peggy Sue™, or Sally Sue™
- 12–230 kDa Separation Module for Jess, Abby, or Wes (PN SM-W004)
- 12–230 kDa Separation Module for Peggy Sue or Sally Sue (PN SM-S001)
- Anti-Rabbit Detection Module for Jess, Wes, Peggy Sue or Sally Sue (PN DM-001)
- RIPA Lysis Buffer (PN 040-483)
- Exosome Standards are available from Novus Biologicals, a Bio-Techne Brand
| Antibody Target | Vendor | Part Number | Dilution Used in this Study |
|---|---|---|---|
| Alix | Novus Biologicals | NBP1-49701 | 1:10 |
| Annexin A2 | Cell Signaling Technology | 8235 | 1:250 |
| Annexin V | Novus Biologicals | NB100-1930 | 1:250 |
| CD63 | Abcam | ab68418 | 1:10 |
| CD9 | Cell Signaling Technology | D8O1A | 1:10 |
| Flotillin-1 | Novus Biologicals | NBP1-79022 | 1:250 |
| HER1/EGFR | Cell Signaling Technology | 2646 | 1:50 |
| HSPA8/HSC70 | Cell Signaling Technology | 8444 | 1:50 |
| TSG101 | Sigma-Aldrich | T5826 | 1:10 |
Table 1. Antibodies against exosome targets validated on Simple Western. All antibodies are sourced from rabbit.
Methods
- For analysis on Simple Western, lyse the exosome standards directly in 1X RIPA on ice for 15 minutes and then proceed with assay setup. Typically, a final exosome-derived lysate sample of 0.2 mg/mL was prepared per the product insert of the Separation Module. If you need exosomes for additional applications, first resuspend the exosomes in deionized water per the instructions provided in the product insert of the exosome standards prior to lysis in RIPA buffer. Additional denaturation conditions should be considered when working with specific classes of proteins. For example, when working with membrane proteins, follow the guidelines in Analyzing Integral Membrane Proteins with Simple Western. This protocol also describes treatment of glycosylated proteins with PNGase F.
- Dilute the antibodies in Antibody Diluent 2.
- Follow the default sample preparation and assay conditions for Simple Western.
Results
We examined three exosome standards derived from human plasma, human serum and A549 cells, which are adenocarcinoma human alveolar basal epithelial cells. Using these standards, we validated antibodies to common exosome targets listed in Table 1. Certain targets, such as the integral membrane protein Flotillin-1, are canonical exosome markers and ubiquitously expressed in most exosomes. As expected, Flotillin-1 was found in all exosome standards we tested (Figure 1A). Conversely, some proteins are specific to the origin of the exosome. For example, Annexin V, found in the plasma membrane, is a marker of apoptosis and anticoagulant protein in the blood. As such, it was found in A549 and plasma exosomes, but not found in serum-derived exosomes (Figure 1B). Likewise, EGFR is specific for A549-derived exosomes and it was not detected in exosomes derived from serum or plasma (Figure 1C). Finally, CD63 (Alix) was detected in all three exosome standard preparations, but had higher overall expression in A549-derived exosomes (Figure 1D).
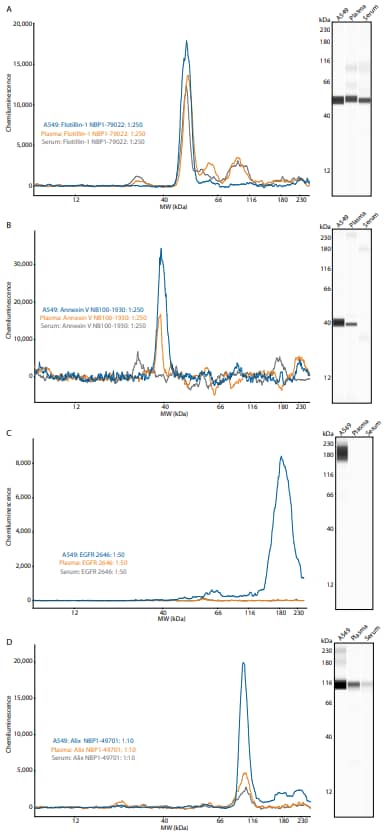
Figure 1. Detection of exosome targets by Simple Western. (A) Flotillin-1 (B) Annexin V (C) EGFR and (D) Alix. Electropherograms are shown (left), as well as lane view (right).
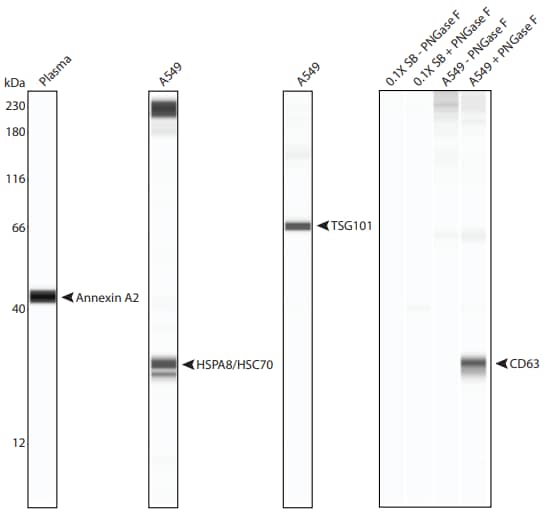
Figure 2. Detection of additional exosome targets by Simple Western. Shown here is the lane view of each target detected on exosome standards from plasma and A549 cells. CD63 gave a signal only upon treatment with PNGase F. The lanes labeled with 0.1X SB contain sample buffer but lack exosome standard. For more information on PNGase F treatment, refer to Analyzing Integral Membrane Proteins with Simple Western.
The other antibodies to common exosome targets listed in Table 1 were validated on Simple Western (Figure 2). Interestingly, a signal for CD63 appeared only upon treatment of the exosome standard with the deglycosylation enzyme PNGase F (Figure 2), which highlights the importance of taking into consideration the specific class of a target protein, for example whether it is glycosylated or membrane-associated.
Conclusion
When reconstituted to a concentration of 1 mg/mL, the exosome standards used here have at least 1x1010 particles/mL. Therefore, at a final sample concentration of 0.2 mg/mL, the theoretical count of exosomes analyzed per well with a 3 µL loading volume is approximately 6 million. However, this is certainly not the lower limit of detecton; we were able to measure a robust signal of Flotillin-1 with as little as 0.03 mg/mL (900,000 particles) of pure exosome sample. Using these standards, we validated antibodies to common exosome targets on Simple Western (Table 1). These results show that Simple Western is an ideal method for analysis of exosomes where sample volume may be of critical importance.
-
Pegtel et al. (2019) Exosomes Annual Review of Biochemistry 88:487–514. PMID: 31220978.
-
Rak (2013) Extracellular Vesicles – Biomarkers and Effectors of the Cellular Interactome in Cancer Frontiers in Pharmacology 4:21. PMID: 23508692.
-
Conlan et al. (2017) Exosomes as Reconfigurable Therapeutic Systems Trends in Molecular Medicine 23:636-650. PMID: 28648185.