Analyzing Integral Membrane Proteins with Simple Western
Introduction
Integral membrane proteins (IMPs) are involved in vital cellular functions of all cells, and account for 20-30% of all open reading frames.1 However, biochemical analysis of IMPs is hindered by their unique properties. Unlike globular proteins, IMPs reside within the lipid-rich hydrophobic membrane that encapsulates cells and subcellular compartments. They often span the membrane multiple times with hydrophobic amino acids clustered in their transmembrane domains. Stripping IMPs from their native environment can bring challenges, and often requires different lysis and denaturation conditions and the addition of detergents.
Before embarking on an experiment to study IMPs, you first need to establish a successful extraction strategy for your IMP of interest. This can include using a lysis buffer rich in detergents and isolating the specific subcellular components through fractionation.2-3 In this protocol, we show you work done by the University of Nevada, where they use ultracentrifugation to fractionate plasma membrane-bound integrins, followed by analysis on Simple Western™. We also highlight some tips for testing sample preparation conditions when working with glycosylated membrane proteins and give recommendations for suitable loading controls for IMPs.
Lysis Considerations for Integral Membrane Proteins
IMPs often require a more stringent lysis procedure for efficient extraction.2,3,6 We recommend using a lysis buffer such as ProteinSimple RIPA Lysis Buffer (PN 040-483) or one that is similarly suitable or established for the extraction of the membrane protein of interest. Check out our Simple Western Buffer Compatibility Guide for a list of tested buffers, including some membrane extraction buffers.
Denaturation Considerations for Integral Membrane Proteins
If specific sample preparation guidance already exists for the target of interest (e.g., from literature), we recommend using those conditions on Simple Western. If no sample preparation guidance exists for your target, we recommend first starting with our default sample preparation conditions. For many membrane proteins, this works well. However some IMPs have the potential to aggregate after being exposed to high heat due to unfavorable refolding of their newly exposed hydrophobic domains.9 For these circumstances, the default Simple Western protocol, like heating a sample at 95 °C for 5 minutes, may not be suitable for your membrane protein and may present no signal at the expected molecular weight (MW) and/or a signal at the upper edge of the separation range used.
If the target aggregates under default sample preparation conditions or you just want to be on the safe side, we recommend setting up a small denaturation test where various heating times and temperatures are assessed. Essentially, make one large master sample that can be split into several tubes. After splitting the sample into individual tubes, treat each tube with a different denaturation condition and run them side by side on your Simple Western instrument to identify the best conditions for the IMP of interest (Figure 1).
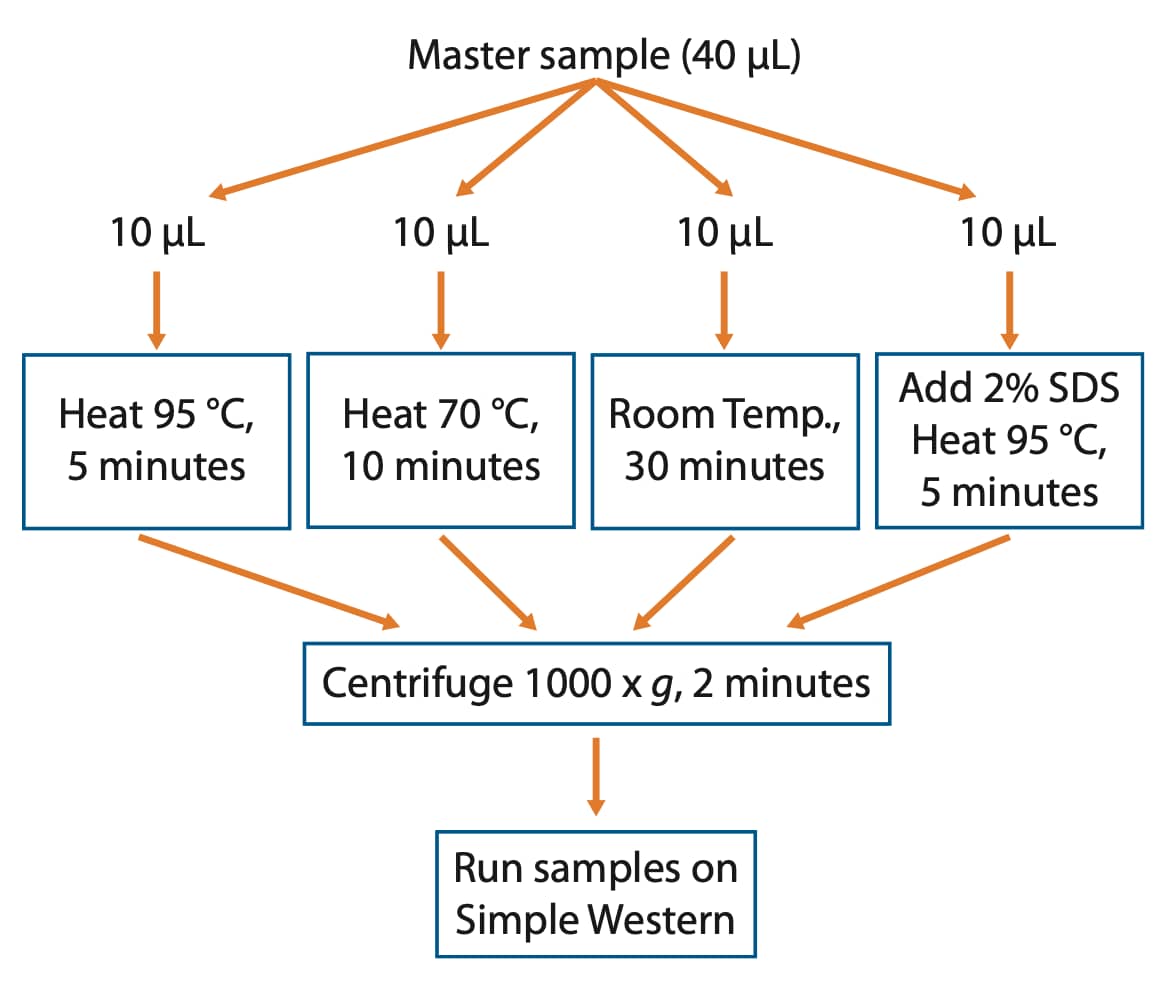
Figure 1. Workflow to identify optimal denaturation conditions for your IMP of interest.
Recommended Denaturation Test for Integral Membrane Proteins
- Make a master sample (40 µL) following the product insert by combining 8 µL 5X Master Mix with 32 µL diluted lysate.
- Split this sample into four tubes, 10 µL per tube.
- Place one tube at 95 °C for 5 minutes, one tube at 70 °C for 10 minutes, and one tube at room temperature for 30 minutes. For the 2% SDS sample, remove 1 µL of the sample and replace with 1 µL of 20% SDS and place the tube at 95 °C for 5 minutes.
- Centrifuge heat-denatured samples for 2 minutes at 1000 x g.
- Run samples in the same Simple Western size run to verify optimal denaturation conditions.
Because IMPs are confined to the boundaries of cells and subcellular compartments, sample preparation may include subcellular fractionation to isolate the specific membrane in which your IMP resides. In the next section, we will show work done by the University of Nevada where isolating membrane fractions is a routine part of their membrane protein analysis.
Fractionation by Ultracentrifugation
The presence of cytosolic signaling machinery can complicate the study of IMPs. Ultracentrifugation is a method to isolate IMPs from the cytosolic proteins.6 Dr. Brian A. Perrino and Ashley Olseen from the Department of Physiology & Cell Biology, University of Nevada Reno School of Medicine, Reno, NV, use ultracentrifugation and the Simple Western platform to study integrin-mediated smooth muscle signaling mechanisms. Integrins are a family of IMPs involved in adhesion-mediated signaling and can form a large array of active dimeric receptors due to the combinatorial pairing of α and β family subunits.4 Receptor combination is restricted (regulated) by the differential expression of integrin subsets in distinct cell types. For example, α8β1 is predominantly expressed in smooth muscle cells, where it has been shown to bind fibronectin and other extracellular matrix proteins.5
Using ultracentrifugation, Dr. Perrino’s lab separated β1-containing integrin dimers into a nearly pure fraction as shown in Figure 2. Analysis of the ultracentrifugation fractions on Simple Western shows that β1 integrin is present in the plasma membrane fractions (100KP) from both human fundus and antrum smooth muscle, and minimally present in the soluble fraction (100KSN), which contains the cytosolic proteins.
The researchers also confirmed the presence of α8 integrin in the same fractions. α8 integrin is present in the 100KP of human stomach smooth muscle (Figure 3, lane 1) but not in skeletal muscle (Figure 3, lane 2). Like β1 integrin, α8 integrin is specifically expressed in smooth muscle and not skeletal muscle.
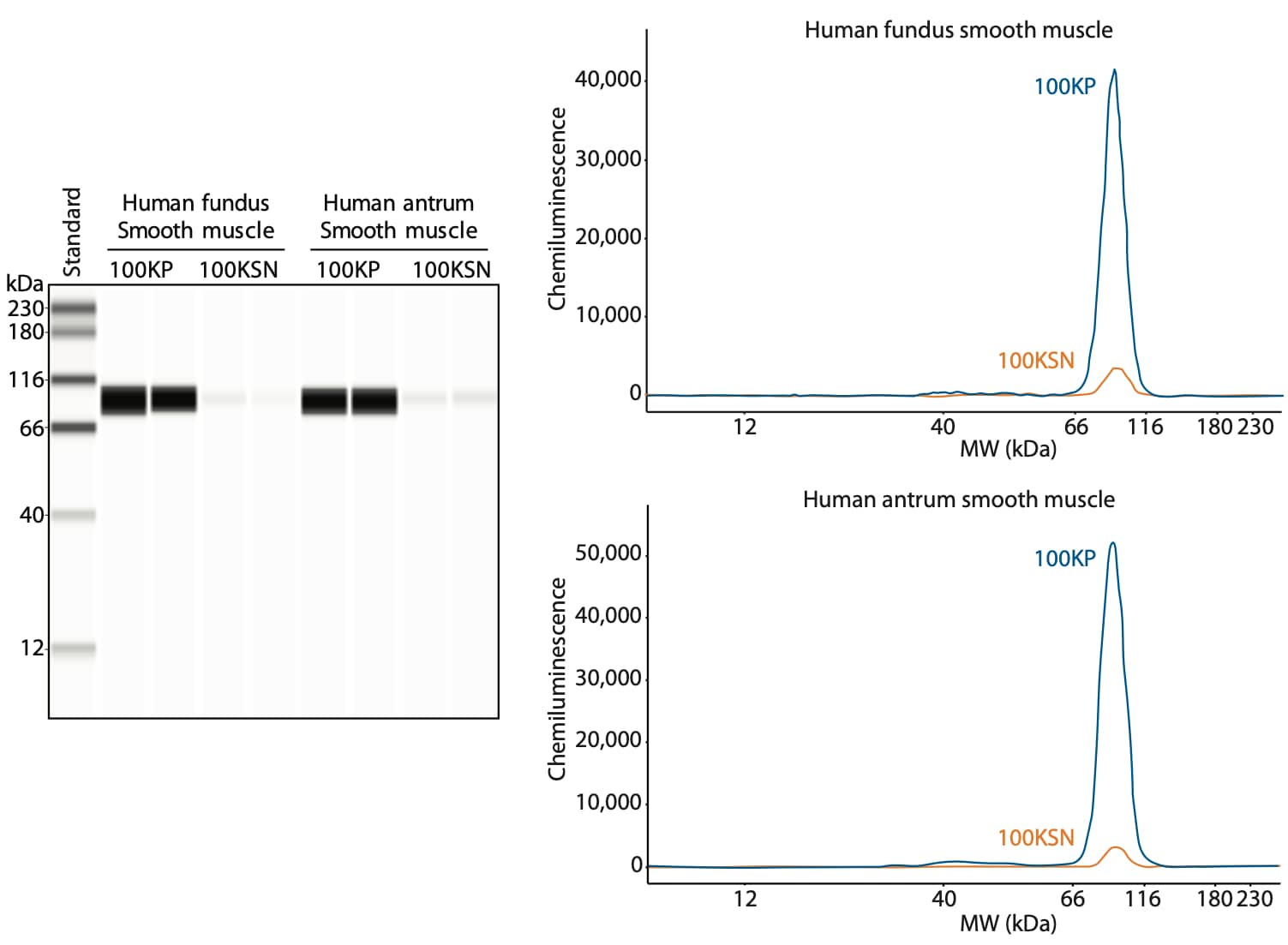
Figure 2. Characterization of β1 integrin expression using Simple Western. Ultracentrifugation fractions were obtained from human stomach smooth muscles from the fundus and antrum by 100,000 x g centrifugation. After centrifugation, the plasma membrane fraction (100KP, blue line) and soluble fraction (100KSN, orange line) were collected. Both fractions were deglycosylated using PNGase F (PNGase F PRIME, Bulldog Bio) and treated with 8 M urea prior to loading 1.5 µg of protein per lane. Samples were loaded in duplicate. β1 integrin was detected using an anti-β1 integrin antibody (Genetex, GTX128839, 1:50).
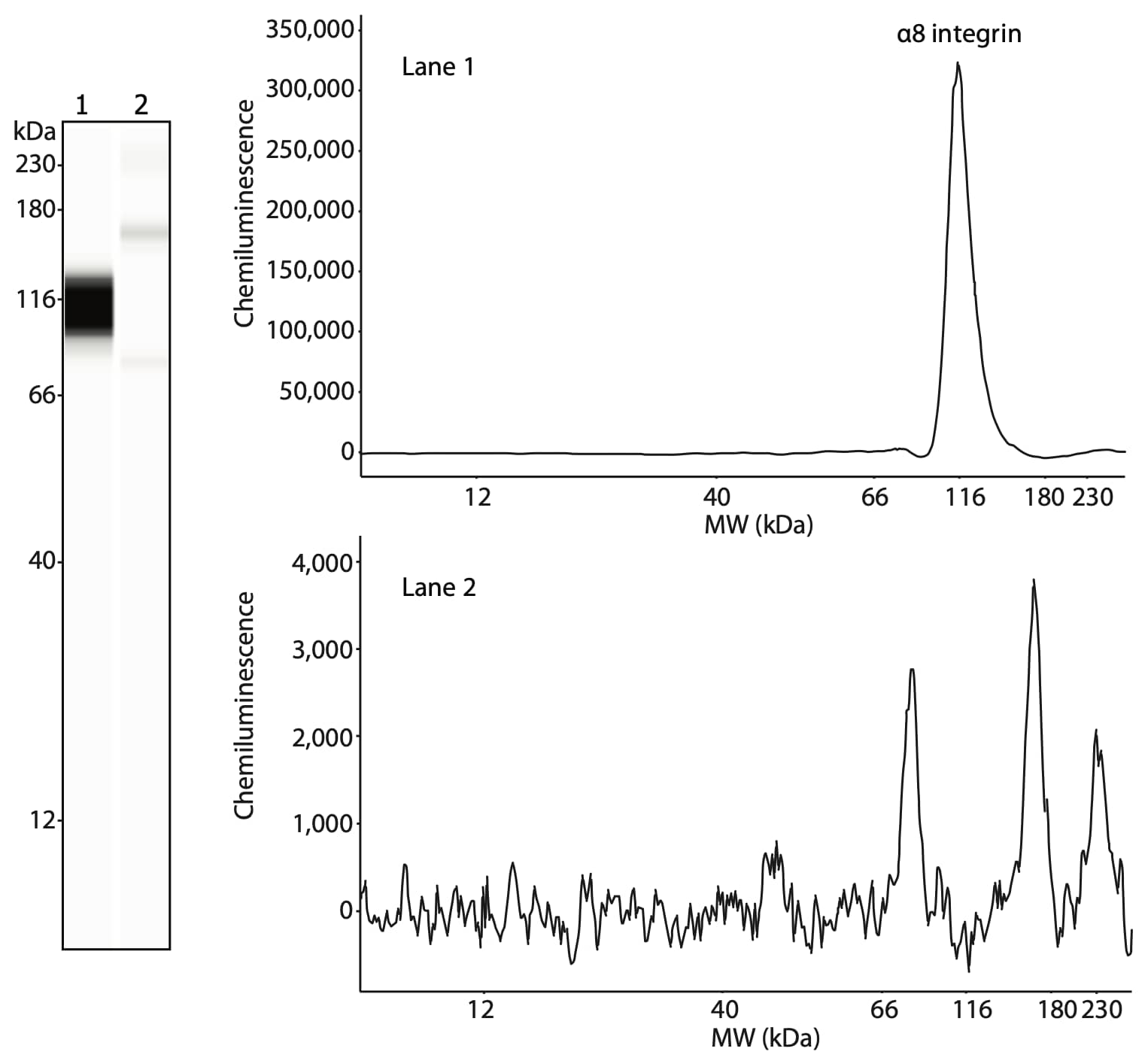
Figure 3. Analysis of α8 integrin, the binding partner of β1 integrin in smooth muscle cells. The plasma membrane fractions from smooth muscle (lane 1) and skeletal muscle (lane 2) were treated with PNGase F prior to loading 1.5 µg of protein per lane. α8 integrin was detected using an anti-α8 integrin antibody (Santa Cruz, sc-365798, 1:50).
Considerations for Glycosylated Proteins
As in traditional Western blot, certain protein features can lead to secondary interactions of the protein of interest with the separation matrix, resulting in shifts of apparent molecular weight compared to the expected MW by amino acid sequence. One common modification that can lead to a MW shift is glycosylation. Here, we describe the protocol developed for and used by Perrino et al. using deglycosylation to verify the glycosylation state of their protein of interest.
Deglycosylation Protocol for Simple Western:
- Perform the deglycoslyation using PNGase F PRIME (Bulldog Bio, NZPP050) under denaturing conditions per the product insert. Another source of PNGase F may be used, but this may require additional optimization.
- We recommend including a control where the enzyme is omitted from the reaction.
- Add 5X Master Mix directly to the control and deglycosylated samples and denature using optimized sample preparation conditions.
- Load the control and deglycosylated samples onto the Simple Western plate.
Note: Dr. Perrino’s group included an additional (and optional) denaturation step by adding 8 M urea to the sample after deglycosylation.
A Loading Control Recommendation for Membrane Protein Preparations
Roddy and colleagues at Merck used the Na+/K+ ATPase as a membrane loading control when looking at liver microsomes for LDLr expression following anacetrapib treatment on Simple Western (reference included in our Publication Spotlight below). This loading control runs at approximately 110 kDa and is expressed on the plasma membrane of a wide variety of cells and tissues.10
Summary
IMPs are an inherently challenging class of proteins to analyze on any platform. With proper solubilization and denaturation conditions, these proteins can easily be resolved and detected by Simple Western. Additional subcellular fractionation, deglycosylation or other treatment may be beneficial or required, as shown for integrins.
Here’s a select list of publications where your colleagues have overcome the challenges of working with membrane proteins...
Publication Spotlight: Membrane Proteins
- Chronic exposure to short chain fatty acids modulates transport and metabolism of microbiome-derived phenolics in human intestinal cells, E Van Rymenant, L Abrankó, S Tumova, C Grootaert, J Van Camp, G Williamson and A Kerimi, J Nutr Biochem, 2017; 39:156–168.
- Incubation of cocaine cue reactivity associates with neuroadaptations in the cortical serotonin (5-HT) 5-HT2C receptor (5-HT2CR) system, S Swinford-Jackson, N Anastasio, R Fox, S Stutz and K Cunningham, Neuroscience, 2016; 324:50–61.
- Overlapping substrate and inhibitor specificity of human and murine ABCG2, J Bakhsheshian, M Hall, R Robey, M Herrmann, J Chen, S Bates and M Gottesman, Drug Metabolism and Disposition, 2013; 41(10):1805–1812.
- Role of Src family kinases in BDNF-mediated suppression of cocaine-seeking and prevention of cocaine-induced ERK, GluN2A, and GluN2B dephosphorylation in the prelimbic cortex, S Barry and J McGinty, Neuropsychopharmacology, 2017; 42(10):1972–1980.
- Effects of anacetrapib on plasma lipids, apolipoproteins and PCSK9 in healthy, lean rhesus macaques, T Roddy, D McLaren, Y Chen, D Xie, K Dunn, A Kulick, D Szeto, G Forrest, K Albanese, M Donnelly, C Gai, A Gewain, H Lederman, K Jensen, X Ai, P Vachal, K Akinsanya, M Cleary, S Previs, H Dansky and D Johns, Eur J Pharmacol, 2014; 740:410–416.
-
Wallin et al. (1998) Genome-wide analysis of integral membrane proteins from eubacterial, archaean, and eukaryotic organisms Protein Science 7:1029–38. PMID: 9568909.
-
Seddon et al. (2004) Membrane proteins, lipids and detergents: not just a soap opera Biochimica et Biophysica Acta 1666:105–17. PMID: 15519311.
-
Smith (2017) Strategies for the purification of membrane proteins Methods in Molecular Biology 1485:389-400.. PMID: 27730565.
-
Hynes (1987) Integrins: a family of cell surface receptors Cell 48:549–54. PMID: 3028640.
-
Schnapp et al. (1995) The human integrin alpha 8 beta 1 functions as a receptor for tenascin, fibronectin, and vitronectin Journal of Biological Chemistry 270:23196–202. PMID: 7559467.
-
Duquesne et al. (2010) Membrane protein solubilization Heterologous expression of membrane proteins. Methods in Molecular Biology (Methods and Protocols) 601:205–17. PMID: 20099148.
-
Kudo et al. (2013) Mfge8 suppresses airway hyperresponsiveness in asthma by regulating smooth muscle contraction Proceedings of the National Academy of Sciences 110:660-5. PMID: 23269839.
-
Magnelli et al. (2011) Identification and characterization of protein glycosylation using specific endo- and exoglycosidases Journal of Visual Experiments 58:e3749. PMID: 22230788.
-
Parton et al. (2011) Aggregation of model membrane proteins, modulated by hydrophobic mismatch, membrane curvature, and protein class Biophysical Journal 101:691–99. PMID: 21806937.
-
Therien et al. (2000) Mechanisms of sodium pump regulation American Journal of Physiology: Cell Physiology 279:C541-66. PMID: 10942705.