Human FGF-19 Antibody
R&D Systems, part of Bio-Techne | Catalog # AF969


Key Product Details
Species Reactivity
Validated:
Cited:
Applications
Validated:
Cited:
Label
Antibody Source
Product Specifications
Immunogen
Phe27-Lys216
Accession # O95750
Specificity
Clonality
Host
Isotype
Endotoxin Level
Scientific Data Images for Human FGF-19 Antibody
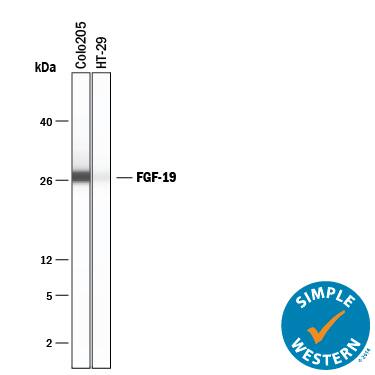
Detection of Human FGF‑19 by Western Blot.
Western blot shows lysates of COLO 205 human colorectal adenocarcinoma cell line, conditioned media from COLO 205 cell line, and HT-29 human colon adenocarcinoma cell line. PVDF membrane was probed with 0.5 µg/mL of Goat Anti-Human FGF-19 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF969) followed by HRP-conjugated Anti-Goat IgG Secondary Antibody (Catalog # HAF017). A specific band was detected for FGF-19 at approximately 22 kDa (as indicated). This experiment was conducted under reducing conditions and using Immunoblot Buffer Group 1.Detection of Human FGF‑19 by Simple WesternTM.
Simple Western lane view shows conditioned media from COLO 205 human colorectal adenocarcinoma cell line and HT-29 human colon adenocarcinoma cell line, loaded at 0.2 mg/mL. A specific band was detected for FGF-19 at approximately 26 kDa (as indicated) using 10 µg/mL of Goat Anti-Human FGF-19 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF969) followed by 1:50 dilution of HRP-conjugated Anti-Goat IgG Secondary Antibody (Catalog # HAF109). This experiment was conducted under reducing conditions and using the 2-40 kDa separation system.Detection of FGF-19 by Western Blot
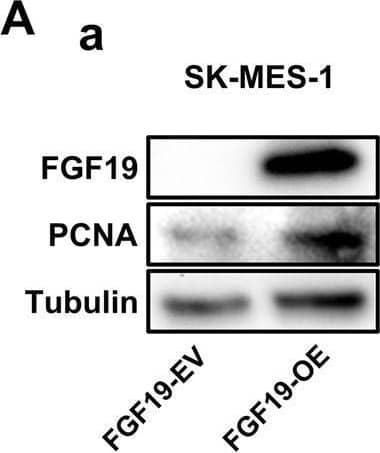
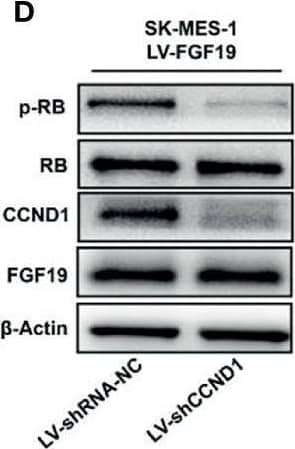
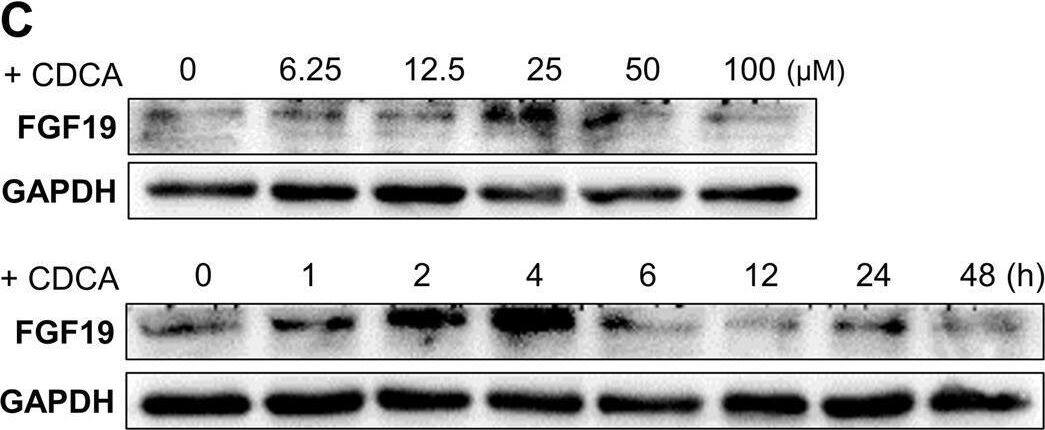
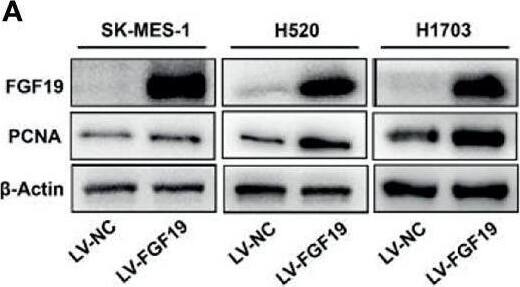
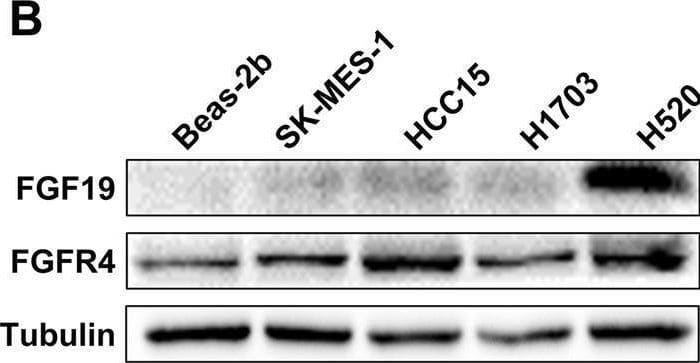
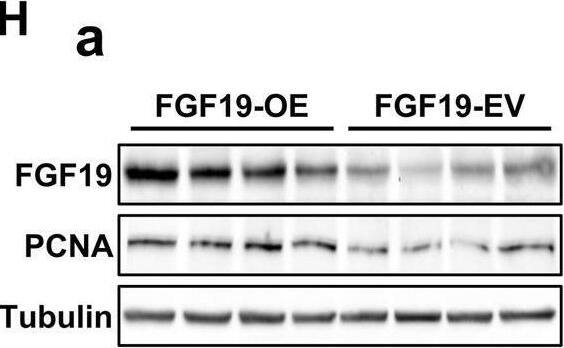
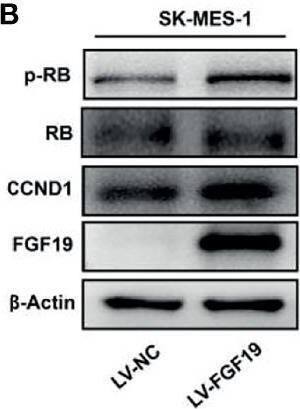
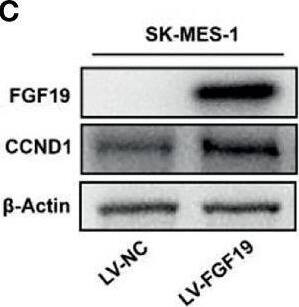
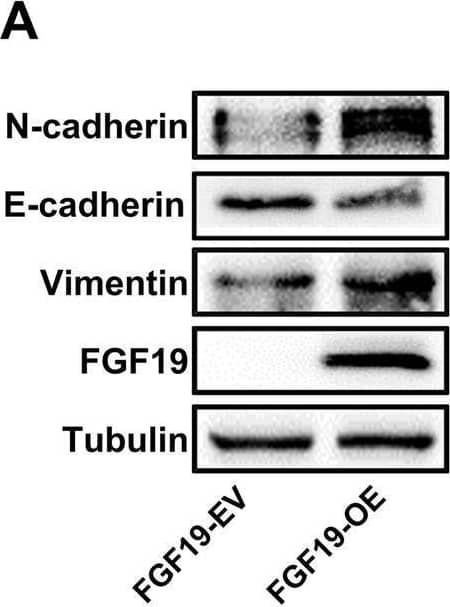
Overexpression of FGF19 promotes the proliferation of LSQ cells in vitro and in vivo.SK-MES-1 and HCC95 cells were transduced with FGF19 overexpression lentivirus (FGF19-OE), or control lentivirus (FGF19-EV) to construct stable cell lines. a Quantification of the overexpression of FGF19 in forms as cellular protein (a), mRNA (b) or secreted protein in conditioned medium (c) in SK-MES-1 cells. b Enrichment plots of PCNA expression signatures according to FGF19 expression levels in an LSQ cohort. c Cell proliferation assays of the SK-MES-1 and HCC95 stable cells with FGF19 overexpression. d Colony formation in SK-MES-1 cells with or without FGF19 overexpression. e The effect of overexpression of FGF19 on the apoptosis induced by cisplatin in HCC95 cell lines. f Enrichment plots of cell cycle signatures according to FGF19 expression levels in an LSQ cohort. g Images of tumor nodules from subcutaneous mouse xenograft model with or without FGF19 overexpression. h (a) Western blot analysis of FGF19 and PCNA expression in tumors; (b) serum samples of FGF19 expression; (c) volume of tumors; and (d) body weight of mice from the two groups. i IHC analysis of Ki-67 expression in tumors. j Oil red O analysis of tumors from the two groups. Data are represented as mean and SEM from three independent experiments. *p < 0.05; **p < 0.01; ***p < 0.001. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/32111983), licensed under a CC-BY license. Not internally tested by R&D Systems.Applications for Human FGF-19 Antibody
Blockade of Receptor-ligand Interaction
Simple Western
Sample: Conditioned media from COLO 205 human colorectal adenocarcinoma cell line and HT‑29 human colon adenocarcinoma cell line
Western Blot
Sample: Conditioned media from COLO 205 cell line and HT‑29 human colon adenocarcinoma cell line
Formulation, Preparation, and Storage
Purification
Reconstitution
Formulation
Shipping
Stability & Storage
- 12 months from date of receipt, -20 to -70 °C as supplied.
- 1 month, 2 to 8 °C under sterile conditions after reconstitution.
- 6 months, -20 to -70 °C under sterile conditions after reconstitution.
Background: FGF-19
Fibroblast growth factor 19 (FGF-19) belongs to the large FGF family which has at least 23 members (1, 2). All FGF family members are heparin-binding growth factors with a core 120 amino acid (aa) FGF domain that allows for a common tertiary structure. FGFs are expressed during embryonic development and in restricted adult tissues. They act on cells of mesodermal and neuroectodermal origin to regulate diverse physiologic functions including angiogenesis, cell growth, pattern formation, embryonic development, metabolic regulation, cell migration, neurotrophic effects and tissue repair (3, 4). Signaling receptors for FGFs are type I transmembrane receptor tyrosine kinases belonging to the Ig superfamily. Four distinct but related classes of FGF receptors, FGF R1, 2, 3, and 4, exist. Through alternative splicing, multiple isoforms for FGF R1, 2 and 3, with distinct ligand recognition profiles, are also generated (4).
Human FGF-19 cDNA predicts a 251 aa precursor protein with a 22 aa signal peptide and a 229 aa secreted mature protein with no potential N-linked glycosylation sites (1, 2). Among FGF family members, human FGF-19 is most closely related to chicken FGF-19 and murine FGF-15, sharing approximately 61% and 51% aa sequence identity, respectively (1, 2, 5). Neither the human orthologue of mouse FGF-15, nor the mouse counterpart of human FGF-19 has been identified. With the exception of adult gall bladder epithelium, FGF-19 expression is restricted to fetal tissues (1, 2). Unlike most FGFs which bind to and activate more than one FGF receptor, FGF-19 is a specific ligand for FGF R4 (2). Similarly, another FGF family member, FGF-7 (KGF), only activates KGF R, the IIIb isoform of FGF R2 (4). During chick embryogenesis, FGF-19 has been shown to act synergistically with Wnt-8c to initiate inner ear development (5).
References
- Nishimura, T. et al. (1999) Biocheim. Biophys. Acta 1444:148.
- Xie, M. et al. (1999) Cytokine 11:729.
- Goldfarb, M. (1996) Cytokine & Growth Factor Reviews 7:311.
- Green, P. et al. (1996) BioEssays 18:639.
- Ladher, R.K. et al. (2000) Science 290:1965.
Long Name
Alternate Names
Entrez Gene IDs
Gene Symbol
UniProt
Additional FGF-19 Products
Product Documents for Human FGF-19 Antibody
Product Specific Notices for Human FGF-19 Antibody
For research use only